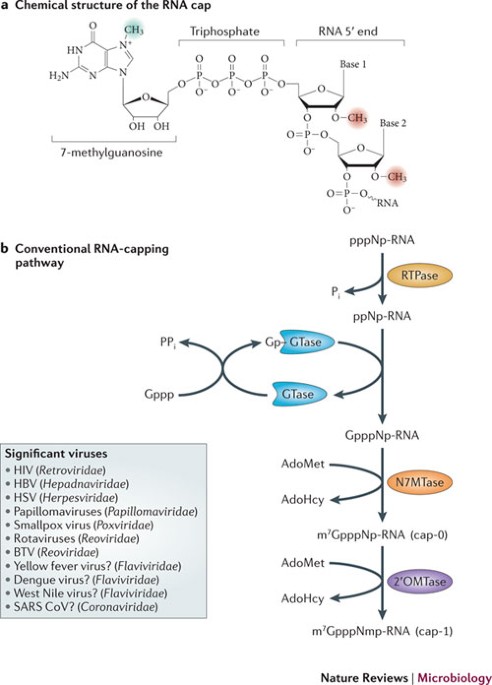
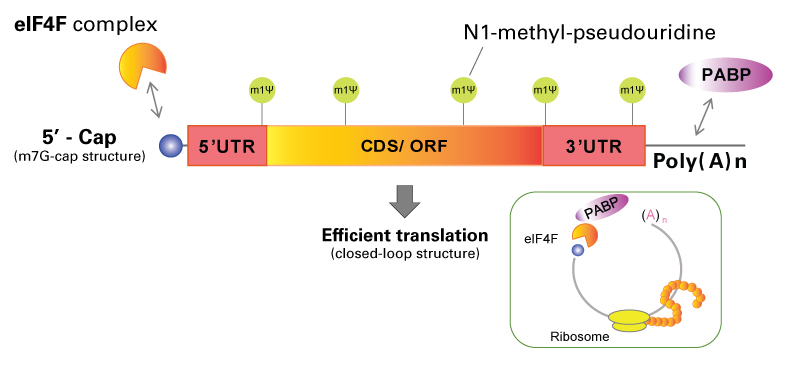
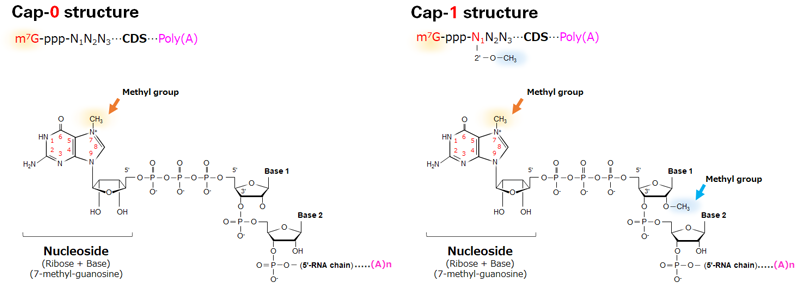
CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology
Rapid Analysis of Synthetic mRNA Cap Structure Using Ion-Pairing RPLC with the BioAccord LC-MS System | Waters